Ribosome profiling (Ribo-Seq)
RNAs can be distinguished by their nucleotide sequences, which confer diverse properties. There are tens to hundreds of thousands of RNA species, and for a long time we had to study them one by one. With the advent of next-generation sequencers, sequences can now be determined in massive parallel (hundreds of millions to billions of reads). This enables determination of RNA identity from sequence information and quantification of RNA abundance from read counts in genome-wide manner (transcriptome analysis).
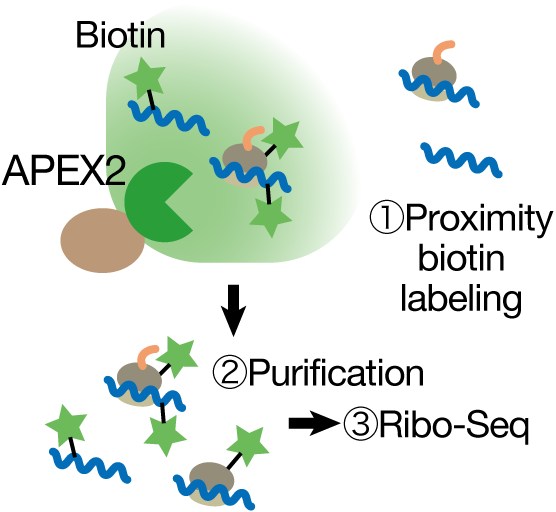
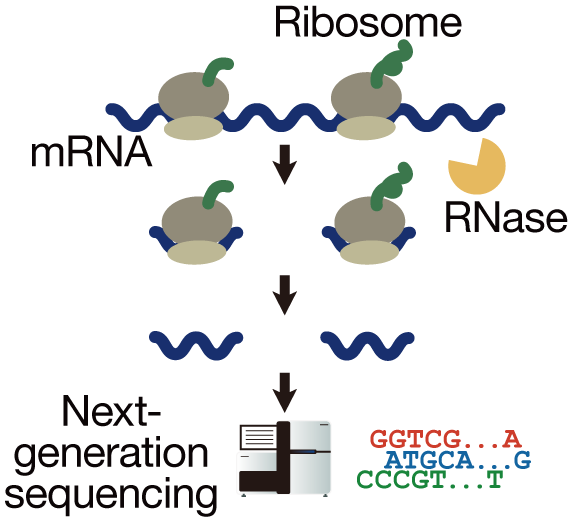
Ribosome profiling (Ribo-Seq) leverages this capability to globally assess translation on mRNAs. In this approach, cell lysates are treated with RNases, which digest most RNA; however, ribosomes protect the region of mRNA they occupy. The protected fragments are sequenced, and the reads are mapped to the genome to identify where and how many ribosomes were bound, i.e., which mRNAs were being translated and to what extent.
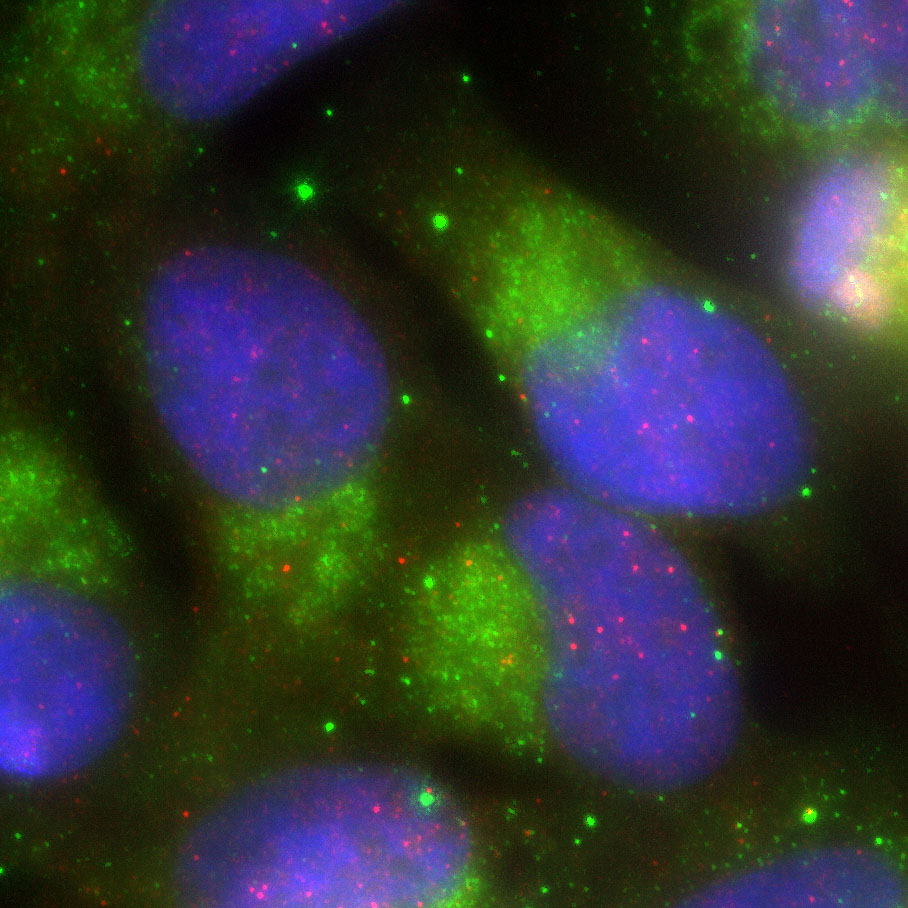
We use Ribo-Seq as a core technology and integrate biochemistry and molecular biology with fluorescence microscopy to reveal how RNA-mediated regulation shapes cellular physiology. We aim for all lab members to become versatile researchers who can perform both wet-lab experiments and dry-computational analyses.